5. Filtering data in 1D#
[ ]:
# %load_ext autoreload
# %autoreload 2
import cmocean
import pandas as pd
import verde as vd
import airbornegeo
[ ]:
data_df = pd.read_csv("data/AGAP_gravity_survey_processed.csv")
data_df = data_df[
[
"easting",
"northing",
"line",
"unixtime",
"grav_disturbance",
"distance_along_line",
]
]
data_df = data_df.sort_values(["line", "unixtime"]).reset_index(drop=True)
data_df.head()
| easting | northing | line | unixtime | grav_disturbance | distance_along_line | |
|---|---|---|---|---|---|---|
| 0 | 1.000024e+06 | 226237.330771 | 1 | 1.229507e+09 | 1186.4 | 0.000000 |
| 1 | 1.000083e+06 | 226246.631269 | 1 | 1.229507e+09 | 342.1 | 59.842447 |
| 2 | 1.000142e+06 | 226255.809132 | 1 | 1.229507e+09 | -1965.9 | 119.693401 |
| 3 | 1.000201e+06 | 226264.969079 | 1 | 1.229507e+09 | 820.0 | 179.545645 |
| 4 | 1.000260e+06 | 226274.156809 | 1 | 1.229507e+09 | 3198.0 | 239.285174 |
5.1. Filter a line#
[7]:
# extract a single line from the survey
line_df = data_df[data_df.line == 4]
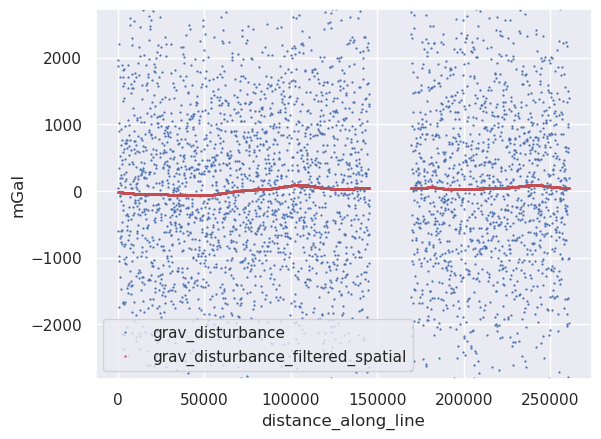
5.1.1. Spatial filtering#
We can perform 1D filtering based on the distance along the flight line.
[10]:
line_df["grav_disturbance_filtered_spatial"] = airbornegeo.filter_line(
line_df,
filter_type="g19000+l", # 19 km low pass gaussian filter
data_column="grav_disturbance",
filter_by_column="distance_along_line",
pad_width_percentage=10,
)
[17]:
ylim = vd.minmax(line_df.grav_disturbance, min_percentile=2, max_percentile=98)
ax = line_df.plot.line(
"distance_along_line",
"grav_disturbance",
style="bp",
ms=0.6,
ylim=ylim,
)
ax = line_df.plot.line(
"distance_along_line",
"grav_disturbance_filtered_spatial",
style="rp",
ms=0.6,
ax=ax,
ylim=ylim,
)
ax.set_ylabel("mGal")
[17]:
Text(0, 0.5, 'mGal')
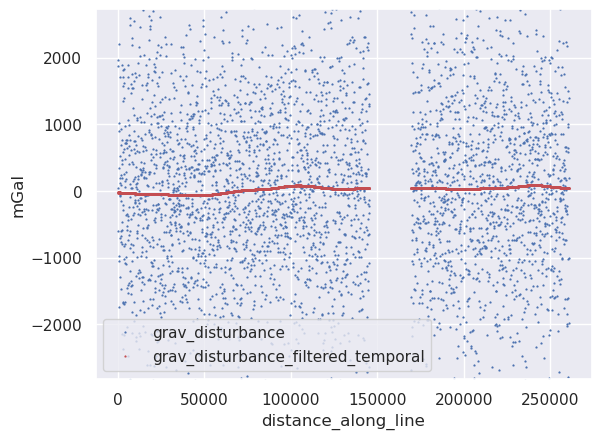
[18]:
line_df = line_df.sort_values("unixtime")
line_df["grav_disturbance_filtered_temporal"] = airbornegeo.filter_line(
line_df,
filter_type="g400+l", # 400 sec low pass gaussian filter
data_column="grav_disturbance",
filter_by_column="unixtime",
pad_width_percentage=10,
)
[19]:
ylim = vd.minmax(line_df.grav_disturbance, min_percentile=2, max_percentile=98)
ax = line_df.plot.line(
"distance_along_line",
"grav_disturbance",
style="bp",
ms=0.6,
ylim=ylim,
)
ax = line_df.plot.line(
"distance_along_line",
"grav_disturbance_filtered_temporal",
style="rp",
ms=0.6,
ax=ax,
ylim=ylim,
)
ax.set_ylabel("mGal")
[19]:
Text(0, 0.5, 'mGal')
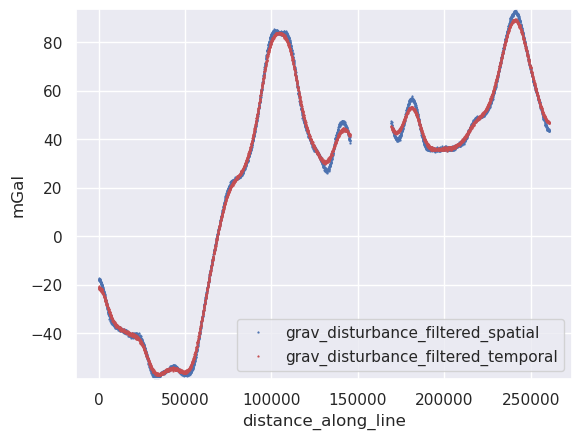
[27]:
ylim = vd.minmax(
line_df[
["grav_disturbance_filtered_spatial", "grav_disturbance_filtered_temporal"]
],
)
ax = line_df.plot.line(
"distance_along_line",
"grav_disturbance_filtered_spatial",
style="bp",
ms=0.6,
ylim=ylim,
)
ax = line_df.plot.line(
"distance_along_line",
"grav_disturbance_filtered_temporal",
style="rp",
ms=0.6,
ax=ax,
ylim=ylim,
)
ax.set_ylabel("mGal")
[27]:
Text(0, 0.5, 'mGal')
5.2. Filter all lines in a survey#
5.2.1. Spatial filtering#
[33]:
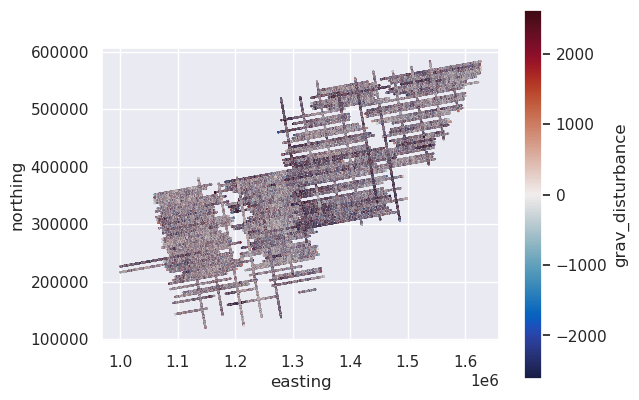
# plot the unfiltered gravity disturbance data
max_abs = vd.maxabs(line_df.grav_disturbance, percentile=95)
ax = data_df.plot.scatter(
"easting",
"northing",
c="grav_disturbance",
s=0.1,
cmap=cmocean.cm.balance,
vmin=-max_abs,
vmax=max_abs,
)
ax.set_aspect("equal")
[34]:
data_df["grav_disturbance_filtered_spatial"] = airbornegeo.filter_line(
data_df,
filter_type="g19000+l", # 19 km low pass gaussian filter
data_column="grav_disturbance",
filter_by_column="distance_along_line",
groupby_column="line",
pad_width_percentage=10,
)
Segments: 100%|██████████| 100/100 [00:03<00:00, 31.87it/s]
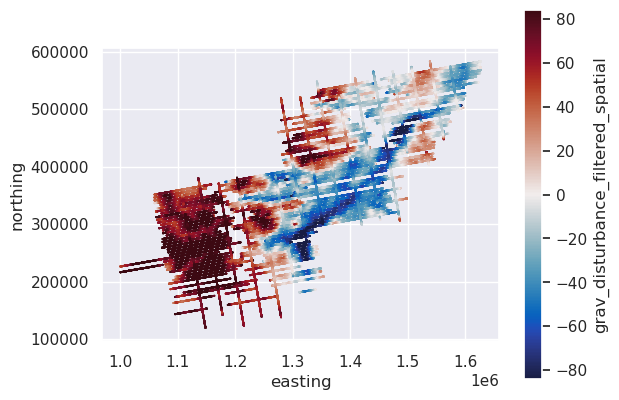
[35]:
# plot the filtered gravity disturbance data
max_abs = vd.maxabs(line_df.grav_disturbance_filtered_spatial, percentile=95)
ax = data_df.plot.scatter(
"easting",
"northing",
c="grav_disturbance_filtered_spatial",
s=0.1,
cmap=cmocean.cm.balance,
vmin=-max_abs,
vmax=max_abs,
)
ax.set_aspect("equal")
5.2.2. Temporal filtering#
[36]:
data_df["grav_disturbance_filtered_temporal"] = airbornegeo.filter_line(
data_df,
filter_type="g500+l", # 500 sec low pass gaussian filter
data_column="grav_disturbance",
filter_by_column="unixtime",
groupby_column="line",
pad_width_percentage=10,
)
Segments: 100%|██████████| 100/100 [00:04<00:00, 23.94it/s]
[37]:
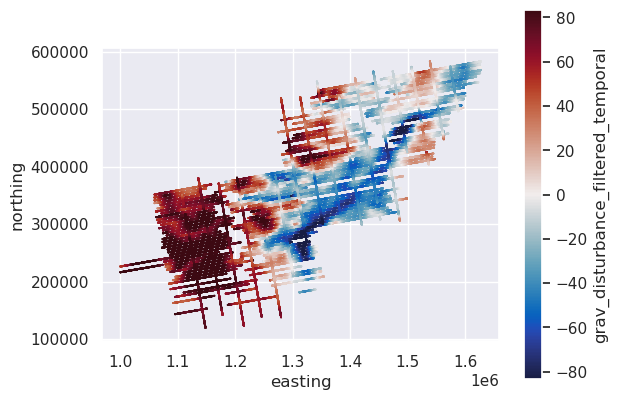
# plot the filtered gravity disturbance data
max_abs = vd.maxabs(line_df.grav_disturbance_filtered_temporal, percentile=95)
ax = data_df.plot.scatter(
"easting",
"northing",
c="grav_disturbance_filtered_temporal",
s=0.1,
cmap=cmocean.cm.balance,
vmin=-max_abs,
vmax=max_abs,
)
ax.set_aspect("equal")